Our aim was to identify and address the bottlenecks for clinical implementation of precision medicine and lay the foundation for an ICT platform to facilitate the development of data driven decision support.
About us
The national lighthouse project BigMed was funded by the Norwegian Research Council and a handful of dedicated partners. The consortium was hosted by Oslo University Hospital and included academic and industrial partners, as well as patient organisations.
BigMed objective
Precision medicine
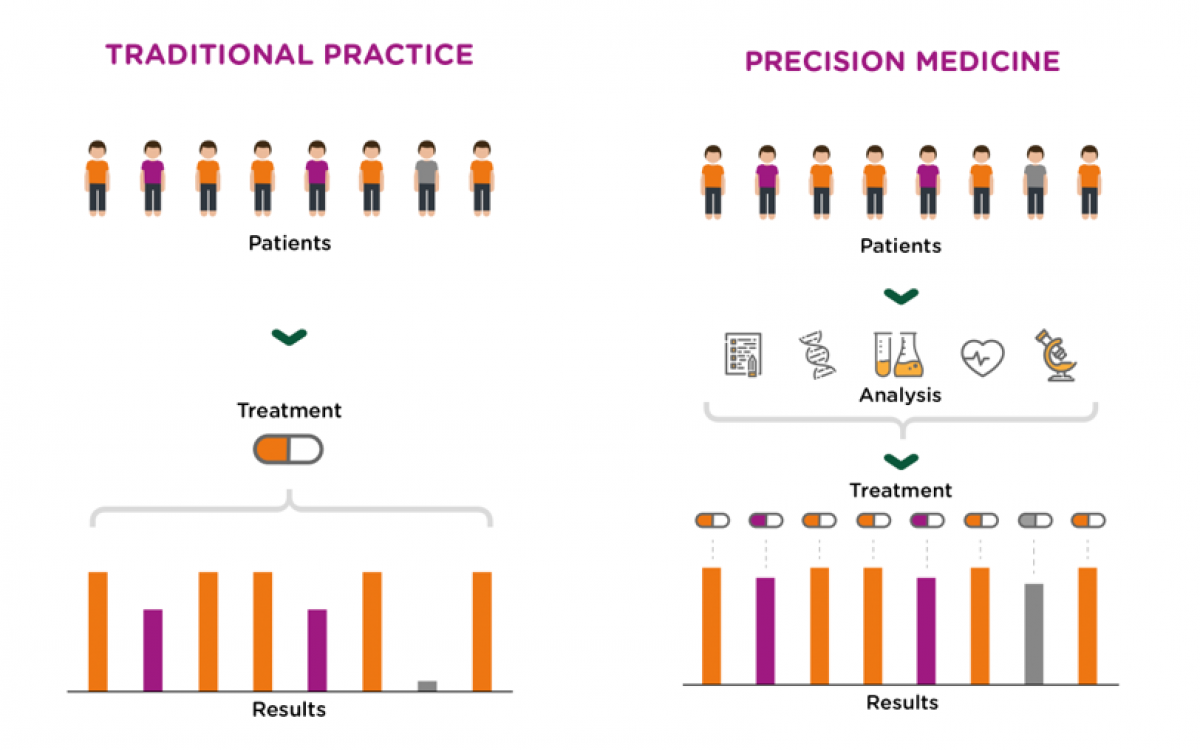
Clinical management of disease is rapidly changing due to developments in diagnostic and therapeutic tools. Traditionally, medical practices have been limited by our inability to obtain sufficient amounts of information for each patient, and a lack of treatment options. This picture is changing as focus shifts from generating relevant data to the challenge of ensuring that all relevant data is available, interpreted and analyzed in a timely manner. Emerging and novel high-throughput technologies can provide large amounts of relevant data and detailed molecular characterization of each patient. Today, decisions on diagnostics and treatment involve cross-disciplinary teams that have to consider large amounts of data. This process can be completely transformed by ICT-systems enabled for structuring, linking and opening for cross-sectional analysis of all available information.
“Can big data analysis improve healthcare services?”
How we work
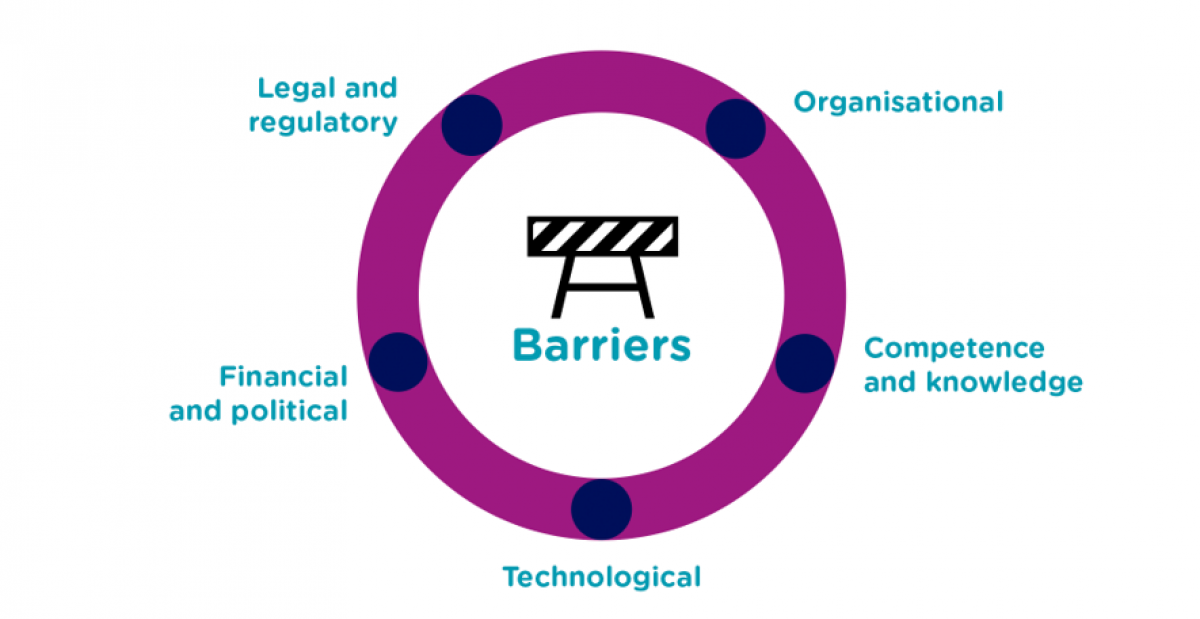
Through developing and demonstrating the implementation of tools and solutions that create value for patients and clinicians, BigMed has gathered knowledge and shared with a wide network of stakeholders in the area of health data. Working in multidisciplinary teams helped tackle the barriers that were identified along the way and finding solutions to help pave the way for precision medicine. The results are documented in our publications and the project articles on this webpage.
“BigMeds main objective has been to identify and address the bottlenecks in precision medicine”
BigMed work packages
The work in BigMed was organized through four clinical areas: rare diseases, colorectal cancer, sudden cardiac death and frost bites.
The suggested solutions were based on the clinical needs identified in each disease area and will focus on the following themes: ICT infrastructure and data provisioning, legal and ethical considerations, bioinformatic pipelines and data sharing.
We are happy to cooperate and share experiences from our work, please get in touch!
“The learning healthcare system of tomorrow has the potential to continuously integrate data and implement new knowledge – meaning immediate benefit to the patients”

